Chemestry: molecules¶
| Author(s): | Joren Retel & Peter Koppatz |
|---|---|
| Tags: | Chemestry, molecules |
Objectives¶
 |
Atoms and molecules – an exiting use case for 3D. With this material it is possible to create molecules, other experiments outside of chemistry and just for fun. A script is guiding you to create molecules or other objects. |
Instructions¶
| Tasks: |
|---|
- Download:
Zipfile with example data (PDB) - Open the blend file and have a look at the examples.
- Add new molecules to the collection.
- Let students construct a molecule from a given set of atoms.
- Working with atoms should be fun, how about constructing a name from a set of atoms.
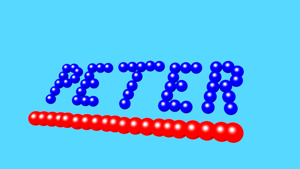

If a real molecule is intergrated in the name, as student can earn some extra points.
| Hint: | All mentioned files are available as a collection within the blend-file. |
|---|
PDB-Format¶
We use the structure of a PDB-File. The documentation about the PDB format is available at:
or the short version in the following table:
| COLUMNS | DATA TYPE | CONTENTS |
|---|---|---|
| 1 - 6 | Record name | “ATOM” |
| 7 - 11 | Integer | Atom serial number. |
| 13 - 16 | Atom | Atom name. |
| 17 | Character | Alternate location indicator. |
| 18 - 20 | Residue name | Residue name. |
| 22 | Character | Chain identifier. |
| 23 - 26 | Integer | Residue sequence number. |
| 27 | AChar | Code for insertion of residues. |
| 31 - 38 | Real(8.3) | Orthogonal coordinates for X in Angstroms. |
| 39 - 46 | Real(8.3) | Orthogonal coordinates for Y in Angstroms. |
| 47 - 54 | Real(8.3) | Orthogonal coordinates for Z in Angstroms. |
| 55 - 60 | Real(6.2) | Occupancy. |
| 61 - 66 | Real(6.2) | Temperature factor (Default = 0.0). |
| 73 - 76 | LString(4) | Segment identifier, left-justified. |
| 77 - 78 | LString(2) | Element symbol, right-justified. |
| 79 - 80 | LString(2) | Charge on the atom. |
Part of a PDB file:
1 2 3 4 5 6 7 8
12345678901234567890123456789012345678901234567890123456789012345678901234567890
ATOM 145 N VAL A 25 32.433 16.336 57.540 1.00 11.92 A1 N
ATOM 146 CA VAL A 25 31.132 16.439 58.160 1.00 11.85 A1 C
ATOM 147 C VAL A 25 30.447 15.105 58.363 1.00 12.34 A1 C
ATOM 148 O VAL A 25 29.520 15.059 59.174 1.00 15.65 A1 O
Simple version or teaching¶
To experiment with this file format we only need the coordinates an the type of atom. So the following simpler structure remains (a water molecule in this example):
########## PDB-Struktur(vereinfacht) ##########
# H20 (Wasser, Water)
ATOM -0.544 -0.257 -0.228 H
ATOM 0.391 0.456 0.228 H
ATOM -0.332 0.44 -0.016 O
Important for the representation is the use of different diameters and colors for each type of atom:
| Atom | Diameter | Color |
|---|---|---|
| Hydrogen | 0,32 | blue |
| Carbon | 0,91 | green |
| Knitted fabric | 0,75 | blue |
| Phospor | 1,06 | red |
| Oxygen | 0,73 | red |
| unknown Atom | 1,0 | black |
Content of the Blender file¶
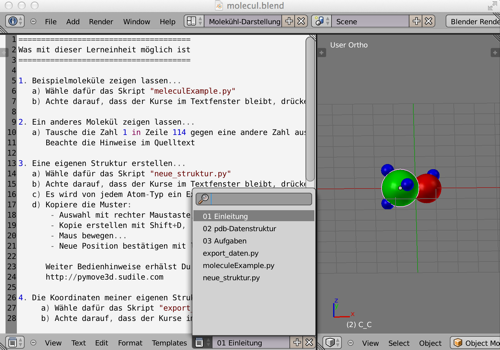
- 01 Introduction (listing of all possibilities)
- 02 PDB data structure (with examples, used by the standard)
- 03 Tasks
- export_data.py (save the coordinates for a new structure to a file)
- moleculeExample.py (Hydrogen, part of a DNA, methanol ...)
- new_struktur.py (a template with one unit of each type of atom)
More informations about this topic¶
A extension for Blender, with more possibilities as in this station shown...
http://wiki.blender.org/index.php/Extensions:2.6/Py/Scripts/Import-Export/PDB
